(Manfred Binder and David S. Hibbett 9/18/2003)
Resuspending freeze-dried oligonucleotides
Oligonucleotides are usually shipped in dry form. The dried DNA pellet becomes dislodged from the bottom of the tube during shipping and it can easily fly out of the tube when first opened, particularly as electrostatic attraction is present. For this reason:
Always briefly centrifuge oligos before opening for the first time.
We dissolve the stock oligo in sterile dH2O which must be freshly autoclaved. Alternatively, TE buffer (10 mM Tris pH 8.0, 1 mM EDTA) can be used.
For convenience, make a freezer stock at 100 µM concentration (which should be thawed infrequently). Adding a volume of dH2O (µL) equal to ten times the number of nanomoles of DNA present in the tube (as noted on the spec sheet provided with the oligo) will produce a stock solution at this concentration. [1 µM = 1 µmole/L or 1 pmole/µL].
For example, dissolve 50 nmoles (= quantity) of oligo in 500 µL dH2O to make a 100 µM stock solution (= concentration). Dilute from this stock 1:10 in dH2O (1 part 100 µM oligo solution, 9 parts dH2O) to make a working solution at 10 µM for use in setting up PCR reactions.
Most PCR reactions use 0.1 - 0.5 µM primer. Addition of 1 µL of the 10 µM primer to a 20 µl PCR reaction (total volume) will result in a final primer concentration of 0.5 µM, or a 10 picomoles quantity of the oligo in a 20 µl volume.
Oligos used in sequencing reactions have lower concentrations at 2 pmoles/µl. For example, use a 10 µM stock and prepare a 1:5 dilution. We use up to 3 picomoles of primer in 12 µl sequencing reactions.
Primer sequences
The conserved rDNA primers that we use for PCR and sequencing have been mostly developed in the Bruns lab and the Vilgalys lab. Visit their web pages for additional information and for a greater choice of primer sequences. The DeepHypha web page provides several primer links and is also summing-up primer sequences for protein coding genes like atp6 (Bruns lab), RPB1 and RPB2 (Hall lab), EF-1a (Steve Rehner), and new primers developed for the AFTOL project, partly including appendant protocolls.
The following is a list of primers currently in use in the Hibbett lab. Check the protocol page for updates on primers of protein-coding regions. Primer sequences (5"—3’), hybridization regions and their relative position are given where applicable. It is also indicated which primers are used for PCR and which primers are used for sequencing (SEQ) purpose.
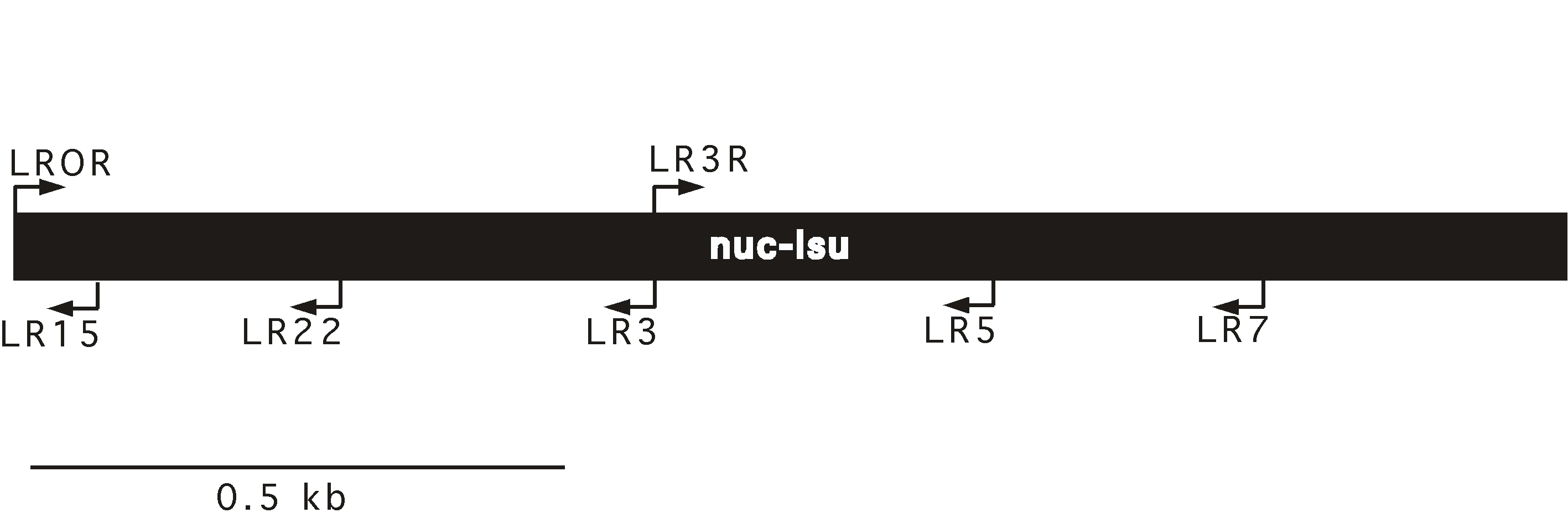
Nuclear large subunit rDNA (nuc-lsu, 25S, 28S)
| Name | Sequence 5’-3’ | Position | Notes |
| LROR | ACC CGC TGA ACT TAA GC | 26-42 | Vilgalys lab |
| LR15 | TAA ATT ACA ACT CGG AC | 154-138 | |
| LR22 | CCT CAC GGT ACT TGT TCG CT | 364-344 | |
| LR3 | CCG TGT TTC AAG ACG GG | 651-635 | |
| LR3R | GTC TTG AAA CAC GGA CC | 638-654 | |
| LR5 | TCC TGA GGG AAA CTT CG | 964-948 | |
| LR7 | TAC TAC CAC CAA GAT CT | 1141-1125 | |
| nuc-lsu primer map
|
|||
PCR: LR0R—LR5 (LR7)
SEQ:?LR0R, LR22, LR3, LR3R, LR5, (LR7)
Internal transcribed spacer region (ITS region, including the 5.8S gene)
| ITS1 | TCC GTA GGT GAA CCT GCG G | 1773-1791 (18S) | White et al. 1990 |
| ITS1-F | CTT GGT CAT TTA GAG GAA GTA A | 1735-1756 (18S) | Gardes & Bruns 1993 |
| ITS2 | GCT GCG TTC TTC ATC GAT GC | 53-34 | White et al. 1990 |
| ITS3 | GCA TCG ATG AAG AAC GCA GC | 34-53 | White et al. 1990 |
| ITS4 | TCC TCC GCT TAT TGA TAT GC | 57-38 (25S) | White et al. 1990 |
| ITS4-B | CAG GAG ACT TGT ACA CGG TCC AG | 194-172 (25S) | Gardes & Bruns 1993 |
| ITS5 | GGA AGT AAA AGT CGT AAC AAG G | 1749-1770 (18S) | White et al. 1990 |
| 5.8S | CGC TGC GTT CTT CAT CG | 54-38 | Vilgalys lab |
| 5.8SR | TCG ATG AAG AAC GCA GCG | 37-54 | Vilgalys lab |
| ITS primer map
|
|||
PCR:? ITS1 (ITS1F, ITS5)—ITS4 (ITS4, LR15, ITS4-B) alternatives in parentheses.
SEQ:? see above. It is recommended using 5.8SR (ITS3) and 5.8S (ITS2) to sequence larger products (> 800 bp).
Nuclear small subunit rDNA (nuc-ssu, 18S)
| PNS1 | CCA AGC TTG AAT TCG TAG TCA TAT GCT TGT C | 1-31 | K. O’Donnell (s. Hibbett 1996) |
| NS19bc | GTT TCT CAG GCT CCC TCT CCG G | 399-378 | Bruns lab |
| NS19b | CCG GAG AGG GAG CCT GAG AAA C | 378-399 | |
| NS41 | CCC GTG TTG AGT CAA ATT A | 1200-1182 | |
| NS51 | GGG GGA GTA TGG TCG CAA GGC | 1108-1128 | |
| NS6 | GCA TCA CAG ACC TGT TAT TGC CTC | 1439-1416 | White et al. 1990 |
| NS7 | GAG GCA ATA ACA GGT CT GTG ATG C | 1416-1439 | |
| NS8 | TCC GCA GGT TCA CCT ACG GA | 1792-1773 | |
| SR1 | ATT ACC GCG GCT GCT | 578-564 | Vilgalys lab |
| SR1c | AGC AGC CGC GGT ATT | 564-578 | |
| SR4 | AAA CCA ACA AAA TAGA A | 838-820 | |
| SR6 | TGT TAC GAC TTT TACT T | 1760-1744 | |
| nuc-ssu primer map
|
|||
PCR:?1) PNS1—NS41 and 2) NS19b—NS8; alternatively use PNS1—NS8.
SEQ:?for product 1) PNS1, NS19bc, NS41 and 2) NS19b, NS51, NS7, NS8. SR primers and NS6 are alternatives for sequencing.
Mitochondrial large subunit rDNA (mt-lsu)
| ML5 | CTC GGC AAA TTA TCC TCA TAA G | White et al. 1990 | |||
| ML6 | CAG TAG AAG CTG CAT AGG GTC | ||||
| MLIN3 | CGA CAC AGG TTC GTA GGT AG | Bruns lab (see web page for intron sites) | |||
| CML7.5 | CCG CCC CAG TCA AAC TGC C | ||||
| mt-lsu primer map
|
|||||
PCR and SEQ: ML5—ML6. MLIN3 and CML7.5 are alternatives
Mitochondrial small subunit rDNA (mt-ssu)
| MS1 | CAG CAG TCA AGA ATA TTA GTC AAT G | White et al. 1990 | |
| MS2 | GCG GAT TAT CGA ATT AAA TAA C | ||
| U1 | TAA TTT TGG TGC CGA TTG AAC G | Bruns lab | |
| CU6 | TGT GGC ACG TCT ATA GCC CA | ||
| mt-ssu primer map
|
|||
PCR and SEQ: MS1—MS2. U1 and CU6 are alternatives.
Mitochondrial ATPase subunit 6, atp6
| ATP6-1 | ATT AAT TSW CCW TTA GAW CAA TT | Kretzer & Bruns 1999 | |
| ATP6-3 | TCT CCT TTA GAA CAA TTT GA | ||
| ATP6-2 | TAA TTC TAN WGC ATC TTT AAT RTA | ||
| ATP6-4 | AAG TAC GAA WAC WTG WGM TTG |
In experimental stage:
| ATP6-5f | WAT RGT WAG AGA WCA AWT AGG | Binder unpublished, Hibbett lab | |
| ATP6-6r | AAC TAA TAR AGG AAC TAA AGC TA | ||
| atp6 primer map
|
|||
PCR and SEQ: ATP6-1 (ATP6-3)—ATP6-2 (ATP6-4) in any combination.
Note: ATP6-5f and ATP6-6r are not yet extensively tested but they work fine as SEQ primers.
Cytochrome oxidase subunit 3, cox3
| COX3-1 | CAT TTA GTA TCG CCT TCA CCA TGG CC | Kretzer & Bruns 1999 | |
| COX3-2 | AAC AAC CAA ACA ACA TCT ACA AAG TG | ||
| cox3 primer map
|
|||
Translation elongation factor 1a , EF-1a
| EF1-526F | GTC GTY GTY ATY GGH CAY GT | 1-20 | Steve Rehner
(DeepHypha web page) |
| EF1-983F | GCY CCY GGH CAY CGT GAY TTY AT | 336-358 | |
| EF1-1577F | CAR GAY GTB TAC AAG ATY GGT GG | 908-930 | |
| EF1-1567R | ACH GTR CCR ATA CCA CCR ATC TT | 942-920 | |
| EF1-1953R | CCR GCR ACR GTR TGT CTC AT | 1519-1490 | |
| EF1-2218R | ATG ACA CCR ACR GCR ACR GTY TG | 1553-1530 | |
| Efcf | ATY GCY GCN GGT ACY GGY GAR TTC GA | 408-433 | |
| Efdf | AAG GAY GGN CAR ACY CGN GAR CAY GC | 447-472 | |
| Efgr | GCA ATG TGG GCR GTR TGR CAR TC | 1311-1289 | |
| Efir | GCR TGY TCN CGR GTY TGN CCR TC | 472-450 | |
| Efjr | TGY TCN CGR GTY TGN CCR TCY TT | 469-447 | |
| EF-1a primer map
|
|||
PCR:?1) 526F—1567R, 2) EF-df—2218R, 3) 983F—1953R
SEQ:?1) 526F, EF-ir, 1567R; 2) EF-df, 1577F,EF-gr, 2218R; 3) 983F, 1953R
Note:?the suggestions made above are the most reliable combinations in our experience to create overlapping sequences.
The whole gene, however, can be amplified in one or two pieces, while additional PCR products occur more frequently.
Laccase
| Lac 1F | AGC AYT GGC AYG GCT TYT TYC | Omon Isikhuemhen, Vilgalys lab, (Pleurotus, Lentinus, Ganoderma) | |
| Lac 3R | AGA CCR TCA CAR TAY TGR GTG G | ||
| Lac 4R | ATA TCG AAG RAT GRC RGA ATT GAT |
Note: PCR conditions and primer combinations are currently being improved, suggestions will follow later.
Manganese dependant peroxidases and Lignin peroxidases (MnP, LiP)
| E2FB | GAC CTS CAG AAG AAC CTG TTC SA | Omon Isikhuemhen, Vilgalys lab, (Pleurotus) | |
| E8R | CGG AGY TGS GTC TCG ATG AAG A |
Note: PCR conditions and primer combinations are currently being improved, suggestions will follow later.
DNA-directed RNA polymerase II subunit 1, RPB1
| RPB1-Af | GAR TGY CCD GGD CAY TTY GG | Hall lab | |
| RPB1-Cf | CCN GCD ATN TCR TTR TCC ATR TA | ||
| RPB1-Df | TAC AAT GCY GAY TTY GAY GG | ||
| RPB1-Dr | TTC ATY TCR TCD CCR TCR AAR TC | ||
| RPB1-Ff | CAY GCD ATG GGD GGD MGD GAR GG | ||
| RPB1-Fr | CCY TCN CKW CCW CCC ATD GCR TG | ||
| RPB1-G1f | TGR AAD GTR TTD AGD GTC ATY TG | ||
| RPB1-G2r | GTC ATY TGD GTD GCD GGY TCD CC | ||
| aRPB1-Br | TCC GCR CCY TCT TCY TTG G | Matheny et al. 2002 | |
| RPB1 primer map
|
|||
Note: PCR conditions and primer combinations are currently being improved, suggestions will follow later.
DNA-directed RNA polymerase II subunit 2, RPB2
| RPB2-3bF | GGW GGW TAY TTY ATY ATY AAT GG | Hall lab, general | |
| RPB2-6F | TGG GGK WTG GTY TGY CCT GC | ||
| RPB2-6R | GCA GGR CAR ACC AWM CCC CA | ||
| RPB2-7F | ATG GGK AAG CAR GCW ATG GG | ||
| RPB2-7R | CCC ATW GCY TGC TTM CCC AT | ||
| RPB2-11aR | GTG WAT YTT RTC RTC MAC C | ||
| RPB2-11bR | CAA TCW CGY TCC ATY TCW CC | ||
| fRPB2-5f | GAY GAY MGW GAT CAY TTY GG | Hall lab, fungal specific | |
| fRPB2-5R | CCR AAR TGA TCW CKR TCR TC | ||
| fRPB2-7cF | ATG GGY AAR CAA GCY ATG GG | ||
| fRPB2-7cR | CCC ATR GCT TGY TTR CCC AT | ||
| fRPB2-11aR | GCR TGG ATC TTR TCR TCS ACC | ||
| bRPB2-3.1F | ATY GCY CAA GAR MGN ATG GC | Hall lab, basidiomycetes specific | |
| bRPB2-6F | TGG GGY ATG GTN TGY CCY GC | ||
| bRPB2-6.3F | GTY ATY GGT GTN TGG ATG GG | ||
| bRPB2-7R | GAY TGR TTR TGR TCR GGG AAV GG | ||
| bRPB2-7.1R | CCC ATR GCY TGY TTM CCC ATD GC | ||
| bRPB2-10.9R | GTR AAS GGY GTG GCR TCY CC |
In experimental stage:
| RPB2-3F1 | AAR GTY YTK ATY GCM CAR GAG CG | Y.W. Lim unpublished, polypore specific, Hibbett lab | |
| RPB2-6F1 | CAC AAY CAN CAY TGG GGW ATG GT | ||
| RPB2-7F1 | ATG GAT ACN ATG GCS AAY AT | ||
| RPB2-6R1 | ACC ATW CCC CAR TGN TGR TTG TG | ||
| RPB2-7R1 | ATR TTG GCC ATN GTR TCC AT | ||
| RPB2-10R1 | ACC CTT YTG MCC RTG ACR AGA | ||
| RPB2 primer map
|
|||
Note: PCR conditions and primer combinations are currently being improved, suggestions will follow later.
See P. Brandon Matheny’s recently posted updates on RPB2.
Literature cited: